At the PICU (Pediatric Intensive Care Unit) of the University Medical Center Utrecht, we have been working the last years to make the data from our PDMS (Patient Data Management System) available for research. This has resulted in a multilayered system with generic possibilities to extract data and transform those data into a convenient flat table format (as described in a previous post).
The system consists of 3 data sources:
- MetaVision Database. The PDMS database (MetaVision) which contains all the clinical patient data
- PICE Registry. An export file from a Dutch national PICE (Pediatric Intensive Care Evaluation) registry. This contains all admission data and diagnoses to perform PIM (Pediatric Index of Mortality) and PRISM (Pediatric RISk of Mortality) calculations
- Definitions. An online Excel spread sheet that contains all definitions which and how data should be extracted. This file can be modified which results in a modified data extraction and PICURED result step.
There are 5 software libraries that are used to extract and load the data to the PICURED Database:
- MetaVision Import. This library retrieves all data per patient from the MetaVision PDMS database.
- PICE Import. This library can import an Excel data extraction from the PICE registry. To calculate the PIM and PRISM estimated mortality is uses:
- PIM & PRISM calculations. A PIM and PRISM calculation library.
- Observations. This is the main library that can read in the online Definitions spreadsheet and will use that to process all data from the MetaVision and PICE import and turns this into a large flat table format.
- PICURED. The software library that ties all these process together and will write the resulting data sets to the PICURED database.
The processes that occur are:
- All data is extracted from the MetaVision database. The following data types are currently supported:
- Basic patient data (including admission history)
- Monitor and data from equipment (ventilators)
- Laboratory data
- Running processes (like iv lines and tracheal tube)
- Microbiology results
- Observations (for example an EMV score)
- Medication
- Fluid balance data
- All data is extracted from the PICE registry. This includes:
- Diagnoses
- Mortality estimates
- Outcome
- Mortality scores are calculated using the PICE data extraction
- The data is combined from the PICE registry and the PDMS database.
- The data is converted to the PICURED (flat table) data format according to
- The definitions data dictionary
- The resulting data set is loaded into the PICURED database.
When loading the data to a table in the PICURED database there are some options to:
- Aggregate data over a period of time. The default time resolution is per minute. By setting a time resolution, data can be aggregate, for example, per hour or per day.
- Data can be anonymized. This means that all identifiable data is recoded:
- Hospital numbers are turned into GUIDS (random letter digit combinations).
- Date time is turned into the number of minutes from the first data point.
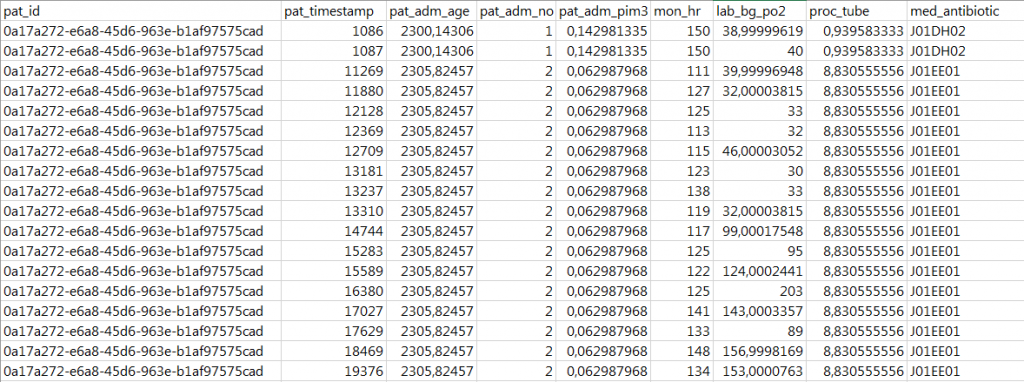
An example data set looks like: